The inspection of 3D z-stack segmentation of cells is enabled by presenting three orthogonal max-intensity projections of 3D cell z-stacks next to three orthogonal max-intensity projections of 3D cell segmentation.
The 3D cell segmentation method follows the method paper and the biological study described in:
One can step through all pairs of cell intensities and cell segmentation results by clicking on the buttons "Next" or "Previous". Labels are assigned to each segmentation by clicking on one of the buttons with the labels under the heading "How would you classify this contact detection?". A user can see already assigned or updated labels by clicking on "Display CVS below". When a user completes the labeling, all assigned labels can be downloaded by clicking on the button "Download CVS".
WARNING: Please, do not reload current web pages during labeling since you would lose all already assigned labels. All labels are kept in the browser cache during a labeling session and therefore switching to a different tab or web page will cause losing the labels.
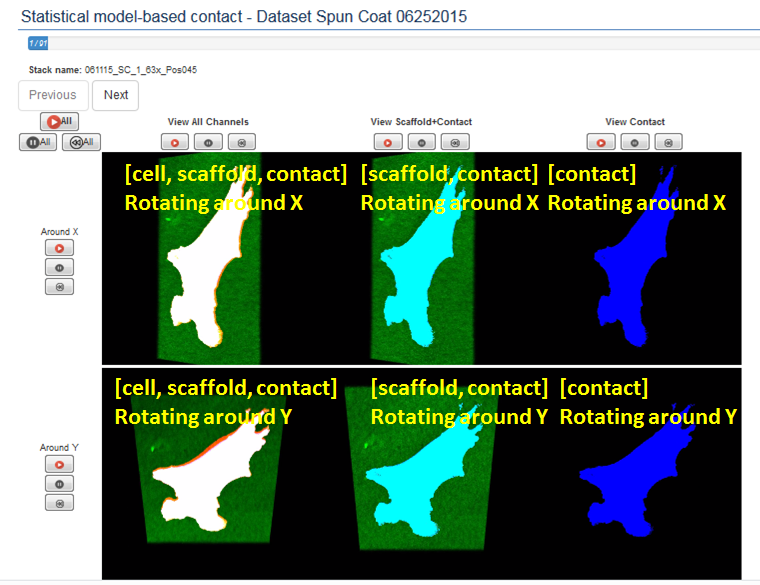
The inspection of 3D cell-contact measurements is enabled by presenting color movies with rotating content around main axes. Specifically, the red-green-blue color is utilized for showing cell (red), scaffold (green) and contacts (blue), and the rotations are around X and Y axes. The arrangement of videos follows a tabular style with two rows corresponding to the two rotations around X (top row) and around Y (bottom row), and three columns. The three columns present [segmented cell, gamma corrected scaffold intensities, detected contacts] information (left column), [gamma corrected scaffold intensities, detected contacts] information (middle column), and [detected contacts] information (right column).

The interpretation of rendered colors for the above assignment is shown in the figure below.
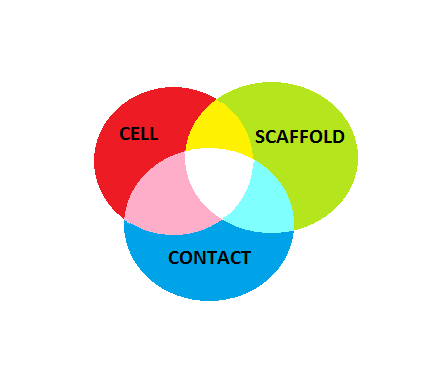
The gamma value for correcting scaffold intensities was set to 1.4 based on visual inspections of gamma corrected zstacks combined with contact points for a range of gamma values between 1.0 and 2.5 (delta gamma = 0.1, two z-stacks, two contact detection methods). The movies corresponding to these settings and used for visual inspection are accessible from the menu bar.
Several related movies can be viewed synchronously or asynchronously, and embedded and original resolutions. Each movie is constructed to last about 9 seconds with 128 frames played at 15 frames per second. All movies are stored as mpeg4 files following the H.264 video codec. Due to the browser support of video players, we recommend viewing the pages in Mozilla Firefox or Google Chrome (see more information about the browser support.)
The cell-scaffold contact points have been detected by 8 methods out of which we selected one statistical and one geometrical model-based method based on orthogonal measurements of single fibers using a scanning electron microscope (SEM). The paper describing the modeling, validation, and verification is work-in-progress.
The labels for each cell-scaffold contact detection span excellent, acceptable and bad.
For the statistical model-based method reporting contact point cloud, the following labels were defined: